NYBSP-1 (ACE2)
Viral Entry & Protein–Protein Interaction Research Peptide
NYBSP-1 is a double hydrocarbon-stapled α-helical peptide derived from Helix-1 of the human angiotensin-converting enzyme 2 (ACE2) receptor. This stapled peptide is designed to stabilize the native ACE2 α-helical conformation involved in binding to the receptor-binding domain (RBD) of the SARS-CoV-2 Spike protein. NYBSP peptides are widely used as research tools to study viral entry mechanisms and protein–protein interactions between viral ligands and host receptors.
Disease Target
Covid 19 SARS COV 2 (block RBD – ACE2 binding).
Stapling Strategy
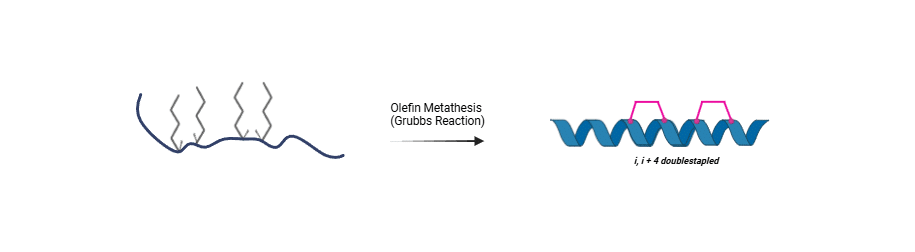
Fmoc-(S)-2-(4-pentenyl)alanine (Fmoc-(S5)-OH, CAS: 288617-73-2) enables the solid-phase peptide synthesis (SPPS) of NYBSP ACE2-derived stapled peptides. During synthesis, Fmoc-(S5)-OH is incorporated at two i,i+4 positions within the ACE2 helix-1 region, followed by ruthenium-catalyzed ring-closing olefin metathesis to generate two (CH₂)₄ hydrocarbon staple. The resulting double stapled peptide blocks SARS-CoV-2 Spike RBD binding (IC₅₀ ≈ 130 nM) and inhibits pseudovirus infection by >90% at 10 μM.
Technical specification
 |
Sequency : | Ac-TIEEQAKT(S5)LDK(S5)NHEAEDLFYQ(S5)SLA(S5)WN-NH2 |
 |
MW : |
3664.08 g/mol |
 |
Purity : | > 95% |
 |
Counter-Ion : | TFA Salts |
 |
Delivery format : | Lyophilized |
Price
| Product | Size | Price € |
Price $ |
| SB326-1mg | 1 mg | 762 | 915 |
| SB326-5mg | 5 mg | 964 | 1157 |
| SB326-10mg | 10 mg | 1291 | 1550 |
For more information about stapled peptides, please visit our dedicated page.
Explore our Stapled Peptide Catalog products. Click HERE to discover additional reference compounds.
Custom Stapled Peptide Services
If your target is not listed, we offer custom stapled peptide design and synthesis. Whether you require sequence optimization, specific modifications, or larger production quantities, our team can support your project. Submit your project details and we will provide a personalized proposal.
References
11 December 2020 - https://doi.org/10.1128/mbio.02451-20
Stapled Peptides Based on Human Angiotensin-Converting Enzyme 2 (ACE2) Potently Inhibit SARS-CoV-2 Infection In Vitro
Abstract
SARS-CoV-2 uses human angiotensin-converting enzyme 2 (ACE2) as the primary receptor to enter host cells and initiate the infection. The critical binding region of ACE2 is an ∼30-amino-acid (aa)-long helix. Here, we report the design of four stapled peptides based on the ACE2 helix, which is expected to bind to SARS-CoV-2 and prevent the binding of the virus to the ACE2 receptor and disrupt the infection. All stapled peptides showed high helical contents (50 to 94% helicity). In contrast, the linear control peptide NYBSP-C showed no helicity (19%). We have evaluated the peptides in a pseudovirus-based single-cycle assay in HT1080/ACE2 cells and human lung cell line A549/ACE2, overexpressing ACE2. Three of the four stapled peptides showed potent antiviral activity in HT1080/ACE2 (50% inhibitory concentration IC50: 1.9 to 4.1 μM) and A549/ACE2 (IC50: 2.2 to 2.8 μM) cells. The linear peptide NYBSP-C and the double-stapled peptide StRIP16, used as controls, showed no antiviral activity. Most significantly, none of the stapled peptides show any cytotoxicity at the highest dose tested. We also evaluated the antiviral activity of the peptides by infecting Vero E6 cells with the replication-competent authentic SARS-CoV-2 (US_WA-1/2020). NYBSP-1 was the most efficient, preventing the complete formation of cytopathic effects (CPEs) at an IC100 of 17.2 μM. NYBSP-2 and NYBSP-4 also prevented the formation of the virus-induced CPE with an IC100 of about 33 μM. We determined the proteolytic stability of one of the most active stapled peptides, NYBSP-4, in human plasma, which showed a half-life (T1/2) of >289 min.
2023 Jan 17;249:115118. doi: 10.1016/j.ejmech.2023.115118
Identification of a short ACE2-derived stapled peptide targeting the SARS-CoV-2 spike protein
Abstract
The design and synthesis of a series of peptide derivatives based on a short ACE2 α-helix 1 epitope and subsequent [i - i+4] stapling of the secondary structure resulted in the identification of a 9-mer peptide capable to compete with recombinant ACE2 towards Spike RBD in the micromolar range. Specifically, SARS-CoV-2 Spike inhibitor screening based on colorimetric ELISA assay and structural studies by circular dichroism showed the ring-closing metathesis cyclization being capable to stabilize the helical structure of the 9-mer 34HEAEDLFYQ42 epitope better than the triazole stapling via click chemistry. MD simulations showed the stapled peptide being able not only to bind the Spike RBD, sterically interfering with ACE2, but also showing higher affinity to the target as compared to parent epitope.
