Asteraceae pPNA
Asteraceae Chloroplast 16S rRNA Blocker
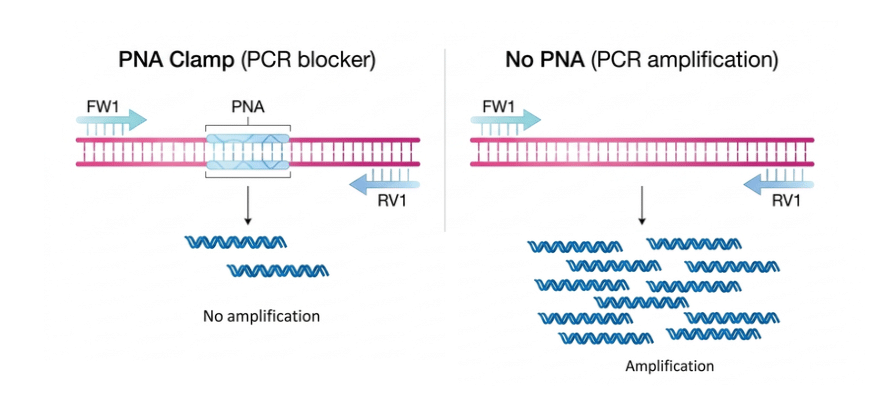
The Asteraceae pPNA is a specialized blocking oligonucleotide designed to suppress the amplification of chloroplast 16S rRNA in Asteraceae plant species. By targeting the conserved ggctcaaccctggacag sequence, this PNA probe significantly reduces host background noise. This enables researchers to bypass abundant chloroplast interference and achieve high-sensitivity detection of rare nuclear targets or microbial DNA.
Main characteristics
Asteraceae pPNA is designed for the targeted suppression of 16S rRNA sequences specifically within the Asteraceae family, allowing a more focused and accurate analysis. Its PNA backbone provides high thermal stability and stronger binding affinity compared to standard DNA, ensuring efficient and reliable blocking. By reducing the amplification of abundant organellar sequences, it improves the detection of low-copy nuclear genes, especially in tissues rich in chloroplast DNA. It is fully compatible with common molecular workflows, including qPCR, RT-PCR, and NGS library preparation, making it easy to integrate into existing protocols. Overall, it enhances sequencing depth and data quality by limiting non-target DNA interference. A terminal glycine can be included in the PNA structure to improve flexibility and reduce steric hindrance, helping to optimize binding without affecting specificity.
Applications
- Reduces chloroplast rRNA amplification in Asteraceae samples for clearer nuclear DNA analysis
- Improves genomic profiling in complex plant tissues by limiting organellar interference
- Supports agricultural and botanical research, including herbicide resistance studies
- Enhances the quality of amplicon sequencing and NGS data
- Minimizes multi-organelle interference in advanced sequencing workflows
Technical specification
 |
Sequency : | ggctcaactctggacag-G |
 |
MW : | 4953.69 g/mol |
 |
Purity : | > 95% |
 |
Counter-Ion : | TFA Salts |
 |
Delivery format : | Lyophilized |
Price
| Product | Size | Price € |
Price $ |
| SB327 - 25nmol | 25 nmol | 399 | 479 |
| SB327 - 50 nmol | 50 nmol | 644 | 772 |
For more information about PNA, please visit our dedicated page.
Explore our PNA Catalog products. Click HERE to discover additional reference compounds.
Custom PNA Services
If your target is not listed, we offer custom PNA design and synthesis. Whether you require sequence optimization, specific modifications, or larger production quantities, our team can support your project. Submit your project details and we will provide a personalized proposal.
References
2018 Aug 18;6(1):144. doi: 10.1186/s40168-018-0534-0.
Chloroplast sequence variation and the efficacy of peptide nucleic acids for blocking host amplification in plant microbiome studies
Abstract
Background: The ability to efficiently characterize microbial communities from host individuals can be limited by co-amplification of host organellar sequences (mitochondrial and/or plastid), which share a common ancestor and thus sequence similarity with extant bacterial lineages. One promising approach is the use of sequence-specific peptide nucleic acid (PNA) clamps, which bind to, and block amplification of, host-derived DNA. Universal PNA clamps have been proposed to block host plant-derived mitochondrial (mPNA) and plastid (pPNA) sequences at the V4 16S rRNA locus, but their efficacy across a wide range of host plant species has not been experimentally tested.
Results: Using the universal PNA clamps, we amplified and sequenced root microbial communities from replicate individuals of 32 plant species with a most recent common ancestor inferred at 140 MYA. We found the average rate of host plastid contamination across plant species was 23%, however, particular lineages exhibited much higher rates (62-94%), with the highest levels of contamination occurring in the Asteraceae. We investigated chloroplast sequence variation at the V4 locus across 500 land plant species (Embryophyta) and found six lineages with mismatches between plastid and the universal pPNA sequence, including all species within the Asteraceae. Using a modified pPNA for the Asteraceae sequence, we found (1) host contamination in Asteraceae species was reduced from 65 to 23%; and (2) host contamination in non-Asteraceae species was increased from 12 to 69%. These results demonstrate that even single nucleotide mismatches can lead to drastic reductions in pPNA efficacy in blocking host amplification. Importantly, we found that pPNA type (universal or modified) had no effect on the detection of individual bacterial taxa, or estimates of within and between sample bacterial diversity, suggesting that our modification did not introduce bias against particular bacterial lineages.
Conclusions: When high similarity exists between host organellar DNA and PCR target sequences, PNA clamps are an important molecular tool to reduce host contamination during amplification. Here, we provide a validated framework to modify universal PNA clamps to accommodate host variation in organellar sequences.
2018 May 2;(135):57561. doi: 10.3791/57561
Exploring the Root Microbiome: Extracting Bacterial Community Data from the Soil, Rhizosphere, and Root Endosphere
Abstract
The intimate interaction between plant host and associated microorganisms is crucial in determining plant fitness, and can foster improved tolerance to abiotic stresses and diseases. As the plant microbiome can be highly complex, low-cost, high-throughput methods such as amplicon-based sequencing of the 16S rRNA gene are often preferred for characterizing its microbial composition and diversity. However, the selection of appropriate methodology when conducting such experiments is critical for reducing biases that can make analysis and comparisons between samples and studies difficult. This protocol describes in detail a standardized methodology for the collection and extraction of DNA from soil, rhizosphere, and root samples. Additionally, we highlight a well-established 16S rRNA amplicon sequencing pipeline that allows for the exploration of the composition of bacterial communities in these samples, and can easily be adapted for other marker genes. This pipeline has been validated for a variety of plant species, including sorghum, maize, wheat, strawberry, and agave, and can help overcome issues associated with the contamination from plant organelles.
Research Article 6 March 2019-https://doi.org/10.1128/msphere.00088-19
Native and Invading Yellow Starthistle (Centaurea solstitialis) Microbiomes Differ in Composition and Diversity of Bacteria
Abstract
Invasive species could benefit from being introduced to locations with more favorable species interactions, including the loss of enemies, the gain of mutualists, or the simplification of complex interaction networks. Microbiomes are an important source of species interactions with strong fitness effects on multicellular organisms, and these interactions are known to vary across regions. The highly invasive plant yellow starthistle (Centaurea solstitialis) has been shown to experience more favorable microbial interactions in its invasions of the Americas, but the microbiome that must contribute to this variation in interactions is unknown. We sequenced amplicons of 16S rRNA genes to characterize bacterial community compositions in the phyllosphere, ectorhizosphere, and endorhizosphere of yellow starthistle plants from seven invading populations in California, USA, and eight native populations in Europe. We tested for the differentiation of microbiomes by geography, plant compartment, and plant genotype. Bacterial communities differed significantly between native and invading plants within plant compartments, with consistently lower diversity in the microbiome of invading plants. The diversity of bacteria in roots was positively correlated with plant genotype diversity within both ranges, but this relationship did not explain microbiome differences between ranges. Our results reveal that these invading plants are experiencing either a simplified microbial environment or simplified microbial interactions as a result of the dominance of a few taxa within their microbiome. Our findings highlight several alternative hypotheses for the sources of variation that we observe in invader microbiomes and the potential for altered bacterial interactions to facilitate invasion success.
