Chloroplast pPNA
Plastid 16S rRNA PNA PCR blocker
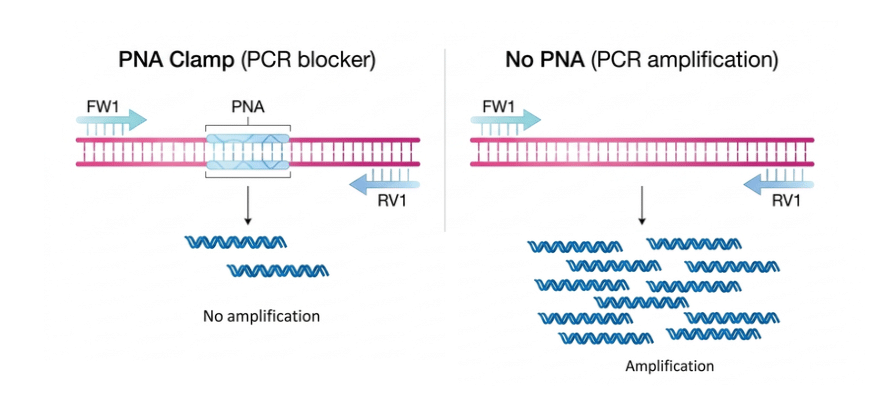
Chloroplast pPNA is a PNA-based PCR blocker designed to specifically bind a conserved chloroplast 16S rRNA sequence in plants (ggctcaaccctggacag), preventing its amplification during PCR and NGS workflows. By reducing interference from chloroplast-derived sequences, it enhances the detection of nuclear genes in complex plant DNA and RNA samples. Thanks to the intrinsic properties of PNA, including high thermal stability, strong sequence specificity, and resistance to enzymatic degradation, pPNA-S is well suited for plant metagenomics, PCR multiplexing, and transcriptomic analyses, especially in samples rich in organelle RNA.
Main characteristics
Chloroplast pPNA PCR blocker specifically targets conserved regions of chloroplast 16S rRNA in plants, helping to limit unwanted amplification during PCR. By reducing chloroplast-derived interference, it improves the amplification and detection of nuclear genes. Its PNA backbone provides strong and selective binding, along with high resistance to enzymatic degradation. It is well suited for plant research and metagenomic applications, and can be easily integrated into both standard and high-throughput PCR workflows. A terminal glycine is added to the PNA structure to improve flexibility and reduce steric hindrance, which can help optimize binding efficiency without affecting specificity.
Applications
- Reduces chloroplast rRNA amplification in plant PCR workflows
- Improves nuclear gene profiling in chloroplast-rich tissues
- Supports RNA-seq and metagenomic enrichment
- Enables multi-target PCR in plant genetics and botany
- Applicable to agricultural genomics and biodiversity studies
- Enhances NGS workflows by depleting organelle DNA
Technical specification
 |
Sequency : | ggctcaaccctggacag-G |
 |
MW : | 4672.42 g/mol |
 |
Purity : | > 95% |
 |
Counter-Ion : | TFA Salts |
 |
Delivery format : | Lyophilized |
Price
| Product | Size | Price € |
Price $ |
| SB337 - 25nmol | 25 nmol | 399 | 479 |
| SB337 - 50 nmol | 50 nmol | 644 | 772 |
For more information about PNA, please visit our dedicated page.
Explore our PNA Catalog products. Click HERE to discover additional reference compounds.
Custom PNA Services
If your target is not listed, we offer custom PNA design and synthesis. Whether you require sequence optimization, specific modifications, or larger production quantities, our team can support your project. Submit your project details and we will provide a personalized proposal.
References
2025 Jan 28;20:14. doi: 10.1186/s40793-025-00674-w
Peptide nucleic acid (PNA) clamps reduce amplification of host chloroplast and mitochondria rRNA gene sequences and increase detected diversity in 16S rRNA gene profiling analysis of oak-associated microbiota
Abstract
Background
Acquiring representative bacterial 16S rRNA gene community profiles in plant microbiome studies can be challenging due to the excessive co-amplification of host chloroplast and mitochondrial rRNA gene sequences that reduce counts of plant-associated bacterial sequences. Peptide Nucleic Acid (PNA) clamps prevent this by blocking PCR primer binding or binding within the amplified region of non-target DNA to stop the function of DNA polymerase. Here, we applied a universal chloroplast (p)PNA clamp and a newly designed mitochondria (m)PNA clamp to minimise host chloroplast and mitochondria amplification in 16S rRNA gene amplicon profiles of leaf, bark and root tissue of two oak species (Quercus robur and Q. petraea).
Results
Adding PNA clamps to PCR led to an overall reduction of host chloroplast and mitochondrial 16S rRNA gene sequences of 79%, 46% and 99% in leaf, bark and root tissues, respectively. This resulted in an average increase in bacterial sequencing reads of 72%, 35%, and 17% in leaf, bark, and root tissue, respectively. Moreover, the bacterial diversity in the leaf and bark increased, with the number of ASVs rising by 105 in the leaf samples and 218 in the bark samples, respectively. In root tissues, where host oak chloroplast and mitochondria contamination were low, alpha and beta diversity did not change, suggesting the PNA clamps did not bias the bacterial community.
Conclusion
In conclusion, this study shows that PNA clamps can effectively reduce host chloroplast and mitochondria PCR amplification and improve assessment of the detected bacterial diversity in Quercus petraea and Quercus robur bacterial 16S rRNA gene sequencing studies.
18 August 2018 Volume 6, article number 144, (2018)
Chloroplast sequence variation and the efficacy of peptide nucleic acids for blocking host amplification in plant microbiome studies
Abstract
Background
The ability to efficiently characterize microbial communities from host individuals can be limited by co-amplification of host organellar sequences (mitochondrial and/or plastid), which share a common ancestor and thus sequence similarity with extant bacterial lineages. One promising approach is the use of sequence-specific peptide nucleic acid (PNA) clamps, which bind to, and block amplification of, host-derived DNA. Universal PNA clamps have been proposed to block host plant-derived mitochondrial (mPNA) and plastid (pPNA) sequences at the V4 16S rRNA locus, but their efficacy across a wide range of host plant species has not been experimentally tested.
Results
Using the universal PNA clamps, we amplified and sequenced root microbial communities from replicate individuals of 32 plant species with a most recent common ancestor inferred at 140 MYA. We found the average rate of host plastid contamination across plant species was 23%, however, particular lineages exhibited much higher rates (62–94%), with the highest levels of contamination occurring in the Asteraceae. We investigated chloroplast sequence variation at the V4 locus across 500 land plant species (Embryophyta) and found six lineages with mismatches between plastid and the universal pPNA sequence, including all species within the Asteraceae. Using a modified pPNA for the Asteraceae sequence, we found (1) host contamination in Asteraceae species was reduced from 65 to 23%; and (2) host contamination in non-Asteraceae species was increased from 12 to 69%. These results demonstrate that even single nucleotide mismatches can lead to drastic reductions in pPNA efficacy in blocking host amplification. Importantly, we found that pPNA type (universal or modified) had no effect on the detection of individual bacterial taxa, or estimates of within and between sample bacterial diversity, suggesting that our modification did not introduce bias against particular bacterial lineages.
Conclusions
When high similarity exists between host organellar DNA and PCR target sequences, PNA clamps are an important molecular tool to reduce host contamination during amplification. Here, we provide a validated framework to modify universal PNA clamps to accommodate host variation in organellar sequences.
Seeds 2025, 4(3), 28; https://doi.org/10.3390/seeds4030028
To Clamp or Not to Clamp: Enhancing Seed Endophyte Metabarcoding Success
Abstract
Seed microbes play crucial roles in plant health, but studying their diversity is challenging due to host DNA contamination. This study aimed to optimise methodologies for investigating seed microbiomes across diverse plant species, focusing on the efficacy of peptide nucleic acid (PNA) clamps to reduce host DNA amplification. We tested PNA clamps on three plant species: Melaleuca quinquenervia (tree), Microlaena stipoides, and Themeda triandra (grasses). The effectiveness of PNA clamps was assessed through in silico analysis, axenic tissue culture, and metabarcoding techniques. In silico analysis confirmed the specificity of PNA clamps to the 16S rRNA gene V4 region of chloroplasts in the grass species. Axenic tissue culture experiments showed that applying PNA clamps at both 1 µM and 0.25 µM concentrations significantly reduced plant DNA amplification. Metabarcoding analyses further confirmed that PNA clamps effectively suppressed host DNA, enhancing microbial diversity estimates across all three species while preserving core microbial taxa. The efficacy of the clamps varied among host species, with T. triandra exhibiting the highest blocking efficacy, and chloroplast clamps outperforming mitochondrial ones. This study demonstrates that PNA clamps are a useful for improving seed endophyte metabarcoding datasets, although they require optimisation for some plant species. This knowledge will contribute to enhancing our understanding of seed microbiome diversity and its ecological implications.
20 December 2016 https://doi.org/10.1111/1755-0998.12645
Identifying the plant-associated microbiome across aquatic and terrestrial environments: the effects of amplification method on taxa discovery
Abstract
Plants in terrestrial and aquatic environments contain a diverse microbiome. Yet, the chloroplast and mitochondria organelles of the plant eukaryotic cell originate from free-living cyanobacteria and Rickettsiales. This represents a challenge for sequencing the plant microbiome with universal primers, as ~99% of 16S rRNA sequences may consist of chloroplast and mitochondrial sequences. Peptide nucleic acid clamps offer a potential solution by blocking amplification of host-associated sequences. We assessed the efficacy of chloroplast and mitochondria-blocking clamps against a range of microbial taxa from soil, freshwater and marine environments. While we found that the mitochondrial blocking clamps appear to be a robust method for assessing animal-associated microbiota, Proteobacterial 16S rRNA binds to the chloroplast-blocking clamp, resulting in a strong sequencing bias against this group. We attribute this bias to a conserved 14-bp sequence in the Proteobacteria that matches the 17-bp chloroplast-blocking clamp sequence. By scanning the Greengenes database, we provide a reference list of nearly 1500 taxa that contain this 14-bp sequence, including 48 families such as the Rhodobacteraceae, Phyllobacteriaceae, Rhizobiaceae, Kiloniellaceae and Caulobacteraceae. To determine where these taxa are found in nature, we mapped this taxa reference list against the Earth Microbiome Project database. These taxa are abundant in a variety of environments, particularly aquatic and semiaquatic freshwater and marine habitats. To facilitate informed decisions on effective use of organelle-blocking clamps, we provide a searchable database of microbial taxa in the Greengenes and Silva databases matching various n-mer oligonucleotides of each PNA sequence.
21 November 2025 Volume 20, article number 148, (2025)
Peptide nucleic acid (PNA) clamps enhance root microbiome profiling in wheat and maize
Abstract
Background
Peptide Nucleic Acid (PNA) clamps represent a crucial molecular tool for reducing host DNA contamination during plant tissue microbiome profiling. This is particularly important when there is sequence similarity between the host organellar DNA (e.g., mitochondrial) and the targeted PCR sequences. However, the effectiveness and optimal concentration of universal PNA clamps can vary between plant species, necessitating a case-by-case evaluation. Here, we assessed the effectiveness of five concentrations (0.0, 0.25, 1.0, 2.0 and 4.0 µM) of mitochondrial and chloroplast PNA blockers (mPNA and pPNA) in reducing the amplification of organellar DNA and enhancing the profiling of prokaryotic communities across root tissues from 34 maize and 27 wheat samples cultivated under various soil and climatic conditions.
Results
We observed that host plant contamination in root samples was consistently high, with an average rate exceeding 95% across all samples. The application of PNA clamps significantly reduced plant host contamination by 2.4–27.2 times in a concentration-dependent manner. This reduction was more pronounced in maize samples than in wheat samples, particularly at lower doses (PNA ≤ 1.0 µM). PNA clamps also increased the read abundance of more than half of the observed microbiome phyla in the root tissues. The most substantial increase in prokaryotic read abundance was observed at a PNA concentration of 1.0 µM, without introducing significant bias to the prokaryotic community.
Conclusions
In conclusion, the introduction of universal PNA clamps during PCR assays significantly reduced amplification of host contamination and enhanced the detection of low-abundance microbiome and the depth of microbial profiling in both maize and wheat root tissues, with effects being concentration- and crop-specific.
