Alpha-Satellite Centromere PNA Probe
Centromere detection by FISH
Alpha-Satellite Centromere PNA Probe is designed to recognize and bind centromeric regions across human chromosomes. Centromeres play a key role in ensuring correct separation of sister chromatids during cell division. This probe specifically targets alpha-satellite DNA, a repetitive sequence characteristic of centromeres, allowing accurate identification of these regions in different experimental applications. The reverse complementary sequence (5' to 3') is aatgcttctgtctagttt.
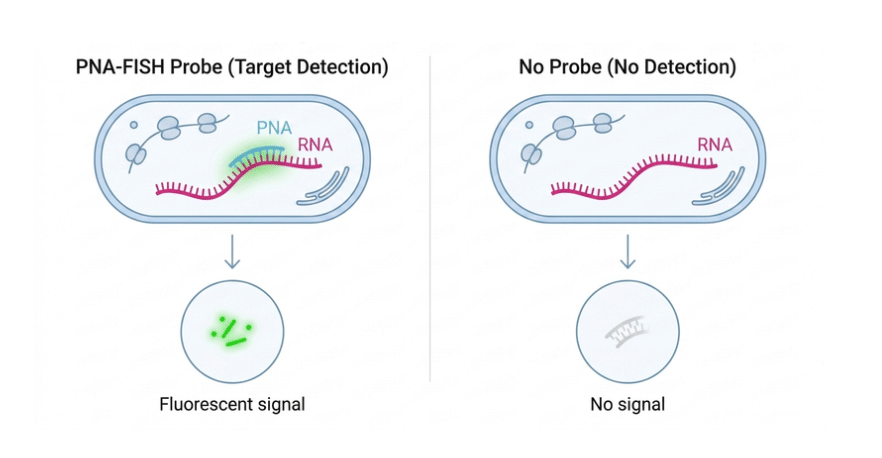
Main characteristics
Alpha-Satellite Centromere PNA Probe is widely applied in Fluorescence In Situ Hybridization (FISH) to detect and visualize centromeric regions. Thanks to its uncharged PNA backbone, it shows stronger and more stable binding to DNA compared to traditional DNA probes, ensuring high specificity. Its ability to recognize centromeres across all human chromosomes makes it a useful tool for a broad range of cytogenetic analyses. The probe is conjugated with FAM (Ex/Em = 495/520 nm), allowing direct detection under fluorescence microscopy. An Ahx spacer (6-aminohexanoic acid, CAS: 60-32-2) is introduced to separate the fluorophore from the recognition sequence, improving accessibility and hybridization efficiency. Additionally, a terminal glycine may be incorporated into the PNA structure to increase molecular flexibility and support optimal binding performance without compromising specificity.
Applications
- Detection of chromosomal abnormalities, including aneuploidies in cancer and genetic disorders
- Chromosome counting and centromere identification in cytogenetic analyses
- Evaluation of genome stability in disease-related research
- Support for karyotyping and chromosomal integrity studies
- Investigation of chromosome segregation defects during cell division
- Useful in both clinical diagnostics and basic cytogenetic research
Technical specification
 |
Sequency : | FAM-Ahx-aaactagacagaagcatt-G |
 |
MW : | 5450.21 g/mol |
 |
Purity : | > 95% |
 |
Counter-Ion : | TFA Salts |
 |
Delivery format : | Lyophilized |
Price
| Product | Size | Price € |
Price $ |
| SB344 - 25nmol | 25 nmol | 516 | 619 |
| SB344 - 50 nmol | 50 nmol | 774 | 929 |
For more information about PNA, please visit our dedicated page.
Explore our PNA Catalog products. Click HERE to discover additional reference compounds.
Custom PNA Services
If your target is not listed, we offer custom PNA design and synthesis. Whether you require sequence optimization, specific modifications, or larger production quantities, our team can support your project. Submit your project details and we will provide a personalized proposal.
References
08 October 2013 Volume 6, article number 42, (2013)
Direct DNA and PNA probe binding to telomeric regions without classical in situ hybridization
Abstract
Background
Fluorescence in situ Hybridization (FISH) utilizes peptide nucleic acid (PNA) probes to identify specific DNA sequences. Traditional techniques have required the heat denaturing of the DNA in formamide followed by multiple hours at moderated temperatures to allow the probe to hybridize to its specific target. Over the past 30 years, advancements in both protocols and probes have made FISH a more reliable technique for both biological research and medical diagnostics, additionally the protocol has been shortened to several minutes. These PNA probes were designed to target and hybridize to both DNA and RNA, and PNA-protein interactions still remain unclear.
Results
In this study we have shown that a telomeric single stranded specific PNA probe is able to bind to its target without heat denaturing of the DNA and without formamide. We have also identified a centromere specific probe, which was found to bind its target with only incubation with formamide.
Conclusions
Certain PNA probes are able to hybridize with their targets with minimal to no denaturing of the DNA itself. This limited denaturing preserves the chromosome structure and may lead to more effective and specific staining.
Volume 70, Issue 3p449-461.e5May 03, 2018
Genome-wide Control of Heterochromatin Replication by the Telomere Capping Protein TRF2
Abstract
Hard-to-replicate regions of chromosomes (e.g., pericentromeres, centromeres, and telomeres) impede replication fork progression, eventually leading, in the event of replication stress, to chromosome fragility, aging, and cancer. Our knowledge of the mechanisms controlling the stability of these regions is essentially limited to telomeres, where fragility is counteracted by the shelterin proteins. Here we show that the shelterin subunit TRF2 ensures progression of the replication fork through pericentromeric heterochromatin, but not centromeric chromatin. In a process involving its N-terminal basic domain, TRF2 binds to pericentromeric Satellite III sequences during S phase, allowing the recruitment of the G-quadruplex-resolving helicase RTEL1 to facilitate fork progression. We also show that TRF2 is required for the stability of other heterochromatic regions localized throughout the genome, paving the way for future research on heterochromatic replication and its relationship with aging and cancer.
Biosensors and Bioelectronics Volume 219, 1 January 2023, 114770
Direct detection of alpha satellite DNA with single-base resolution by using abasic Peptide Nucleic Acids and Fluorescent in situ Hybridization
Abstract
The detection of repetitive sequences with single-base resolution is becoming increasingly important aiming to understand the biological implications of genomic variation in these sequences. However, there is a lack of techniques to experimentally validate sequencing data from repetitive sequences obtained by Next-Generation Sequencing methods, especially in the case of Single-Nucleotide Variations (SNVs). That is one of the reasons why repetitive sequences have been poorly studied and excluded from most genomic studies. Therefore, in addition to sequencing data, there is an urgent need for efficient validation methods of genomic variation in these sequences. Herein we report the development of chemFISH, an alternative method for the detection of SNVs in repetitive sequences. ChemFISH is an innovative method based on dynamic chemistry labelling and abasic Peptide Nucleic Acid (PNA) probes to detect in situ the α-satellite DNA, organized in tandem repeats, with single-base resolution in a direct and rapid reaction. With this approach, we detected by microscopy the α-satellite DNA in a variety of human cell lines, we quantified the detection showing a low coefficient of variation among samples (13.16%–25.33%) and we detected single-base specificity with high sensitivity (82.41%–88.82%). These results indicate that chemFISH can serve as a rapid method to validate previously detected SNVs in sequencing data, as well as to find novel SNVs in repetitive sequences. Furthermore, the versatile chemistry behind chemFISH can lead to develop novel molecular assays for the in situ detection of nucleic acids.
