E. coli 16S-Biotin Probe
E. coli 16S CENPB PNA FISH PROBES
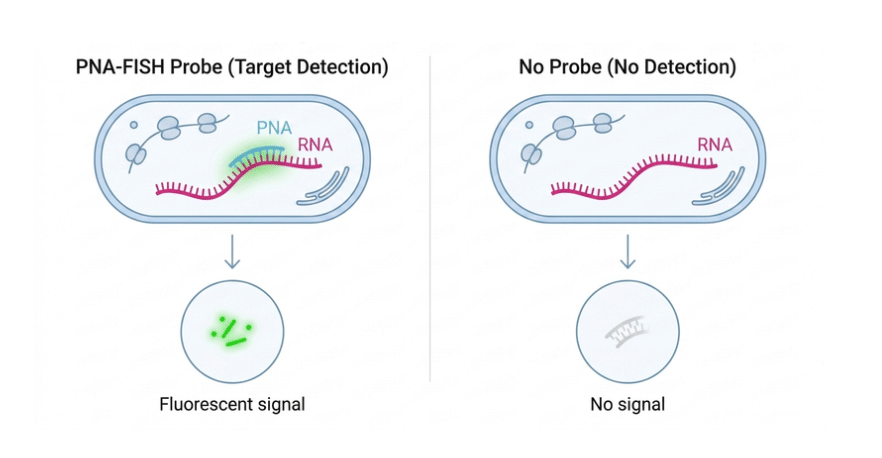
The E. coli 16S-Biotin-Ahx Probe is a sequence-specific PNA probe developed for the selective detection of Escherichia coli, including O157:H7 strains. The probe sequence targets a defined region within the bacterial 16S rRNA, enabling accurate identification. It is designed for use in PNA-FISH, a rapid technique that detects microorganisms directly by hybridizing to intracellular rRNA.
Main characteristics
This PNA probe enables sequence-specific detection of E. coli O157:H7 and is designed for use in PNA-FISH applications targeting rRNA. It provides high specificity for regions associated with enterohemorrhagic E. coli (EHEC) and benefits from the neutral PNA backbone, which ensures strong and stable hybridization. The method allows rapid and sensitive detection without the need for PCR amplification, making it well suited for microbial identification workflows. The probe is conjugated with biotin, enabling detection through streptavidin-based systems for flexible signal development. A terminal glycine can be incorporated to increase flexibility and support optimal binding without affecting specificity.
Applications
- Identification of E. coli O157:H7 in clinical microbiology samples
- Screening for pathogenic E. coli in food safety testing
- Monitoring of environmental samples such as water and agricultural sources
- Use in PNA-FISH methods for microbial detection
- Research on enterohemorrhagic E. coli and similar pathogens
Technical specification
 |
Sequency : | Biotin-caacacacagtgtc-G |
 |
MW : | 4030.94 g/mol |
 |
Purity : | > 95% |
 |
Counter-Ion : | TFA Salts |
 |
Delivery format : | Lyophilized |
Price
| Product | Size | Price € |
Price $ |
| SB346 - 25nmol | 25 nmol | 448 | 537 |
| SB346 - 50 nmol | 50 nmol | 652 | 782 |
For more information about PNA, please visit our dedicated page.
Explore our PNA Catalog products. Click HERE to discover additional reference compounds.
A fluorescent FAM-labeled version of this peptide is available for imaging and uptake studies. Tap the button below to view the FAM-conjugated product.
Custom PNA Services
If your target is not listed, we offer custom PNA design and synthesis. Whether you require sequence optimization, specific modifications, or larger production quantities, our team can support your project. Submit your project details and we will provide a personalized proposal.
References
2013 Oct;79(20):6293–6300. doi: 10.1128/AEM.01009-13
Detection of Escherichia coli O157 by Peptide Nucleic Acid Fluorescence In Situ Hybridization (PNA-FISH) and Comparison to a Standard Culture Method
Abstract
Despite the emergence of non-O157 Shiga toxin-producing Escherichia coli (STEC) infections, E. coli serotype O157 is still the most commonly identified STEC in the world. It causes high morbidity and mortality and has been responsible for a number of outbreaks in many parts of the world. Various methods have been developed to detect this particular serotype, but standard bacteriological methods remain the gold standard. Here, we propose a new peptide nucleic acid fluorescence in situ hybridization (PNA-FISH) method for the rapid detection of E. coli O157. Testing on 54 representative strains showed that the PNA probe is highly sensitive and specific to E. coli O157. The method then was optimized for detection in food samples. Ground beef and unpasteurized milk samples were artificially contaminated with E. coli O157 concentrations ranging from 1 × 10−2 to 1 × 102 CFU per 25 g or ml of food. Samples were then preenriched and analyzed by both the traditional bacteriological method (ISO 16654:2001) and PNA-FISH. The PNA-FISH method performed well in both types of food matrices with a detection limit of 1 CFU/25 g or ml of food samples. Tests on 60 food samples have shown a specificity value of 100% (95% confidence interval [CI], 82.83 to 100), a sensitivity of 97.22% (95% CI, 83.79 to 99.85%), and an accuracy of 98.33% (CI 95%, 83.41 to 99.91%). Results indicate that PNA-FISH performed as well as the traditional culture methods and can reduce the diagnosis time to 1 day.
Volume 48, Issue 1, January 2002, Pages 1-17
PNA for rapid microbiology
Abstract
The acceptance of rRNA sequence diversity as a criterion for phylogenetic discrimination heralds the transition from microbiological identification methods based on phenotypic markers to assays employing molecular techniques. Robust amplification assays and sensitive direct detection methods are rapidly becoming the standard protocols of microbiology laboratories. The emergence of peptide nucleic acid (PNA) from its status as an academic curiosity to that of a promising and powerful molecular tool, coincides with, and complements, the transition to rapid molecular tests. The unique properties of PNA enable the development of assay formats, which go above and beyond the possibilities of DNA probes. PNA probes targeting specific rRNA sequences of yeast and bacteria with clinical, environmental, and industrial value have recently been developed and applied to a variety of rapid assay formats. Some simply incorporate the sensitivity and specificity of PNA probes into traditional methods, such as membrane filtration and microscopic analysis; others involve recent techniques such as real-time and end-point analysis of amplification reactions.
