Globin Reduction PNA Set
Human globin mRNA - PCR Blocker
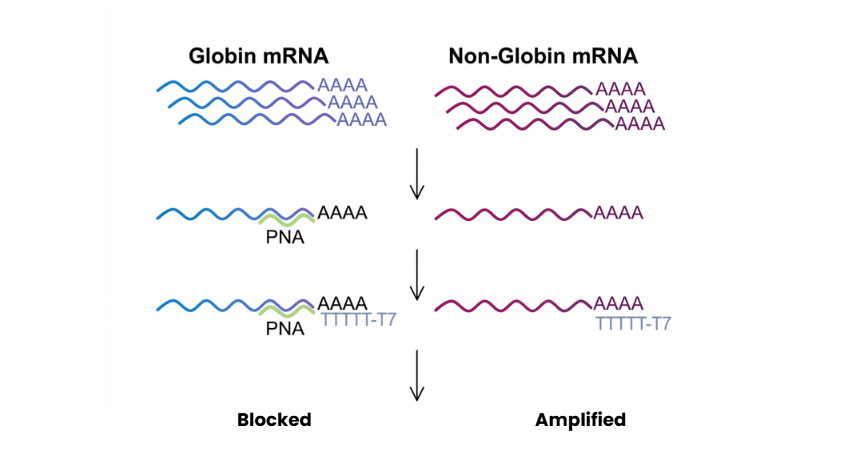
Globin Reduction PNA is a non-enzymatic solution designed to selectively reduce alpha and beta globin mRNA from total RNA extracted from whole blood. It works as a clamp during reverse transcription, binding specifically to globin transcripts and preventing their amplification, which allows more efficient detection of non-globin mRNA targets. Since globin mRNA represents a large proportion of total mRNA in blood (often over 70%), its removal significantly improves the sensitivity and accuracy of downstream gene expression analysis.
Main characteristics
Globin Reduction PNA is designed to limit the impact of globin mRNA during whole blood transcriptome analysis. It works by binding specifically to globin transcripts during reverse transcription, blocking their conversion into cDNA and preventing their amplification in PCR. Since globin mRNA makes up a large portion of total RNA in blood, this reduction step improves the detection of non-globin genes, leading to more sensitive and accurate gene expression analysis.. A terminal glycine is added to the PNA structure to improve flexibility and reduce steric hindrance, which can help optimize binding efficiency without affecting specificity.
Applications
- Reduces globin mRNA interference in whole blood RNA samples
- Improves sensitivity of gene expression analysis in blood transcriptomics
- Enhances RNA-seq data quality by increasing non-globin transcript detection
- Supports biomarker discovery in blood-based studies
- Enables more accurate profiling of low-abundance transcripts
- Applicable to clinical, diagnostic, and research workflows using whole blood RNA
Globin Reduction PNA Set consists of four PNA oligonucleotides specifically designed to target alpha and beta globin mRNA, and is effective for both human and mouse samples:
- Globin- Reduction PNA SB339: taacggtatttggag-G
- Globin-Reduction PNA SB340: gtagttggacttagg-G
- Globin-Reduction PNA SB341: gcccttcataatatc-G
- Globin- Reduction PNA SB342: atccagatgctcaag-G
Why Choose the Complete Globin Reduction PNA Set?
In whole blood transcriptomics, hemoglobin is produced by multiple distinct genes that code for different polypeptide chains, primarily alpha and beta globin. Using a single PNA sequence would only inhibit one specific transcript, allowing the remaining globin mRNA types to dominate the sample and consume the majority of your sequencing resources.
By utilizing the complete Globin Reduction PNA Set, you ensure a comprehensive suppression of all major globin variants. This collective approach is essential to effectively lower the total globin cDNA concentration. By neutralizing the entire family of these abundant transcripts simultaneously, you reallocate sequencing depth toward the rest of the transcriptome. This process is the only way to achieve the sensitivity required to identify rare biomarkers and low-abundance nuclear genes that are otherwise obscured by the high concentration of hemoglobin signals.
Technical specification
 |
Sequency : |
|
 |
MW : |
|
 |
Purity : | > 95% |
 |
Counter-Ion : | TFA Salts |
 |
Delivery format : | Lyophilized |
Price
| Product | Size | Price € |
Price $ |
| SB339-342 - 25nmol | 25 nmol | 354 | 438 |
| SB339-342- 50 nmol | 50 nmol | 582 | 699 |
| Product | Size | Price € |
Price $ |
| SB Globin Reduction PNA Set - 25nmol | 25 nmol | 1021 | 1226 |
| SB Globin Reduction PNA Set- 50 nmol | 50 nmol | 1631 | 1957 |
For more information about PNA, please visit our dedicated page.
Explore our PNA Catalog products. Click HERE to discover additional reference compounds.
Custom PNA Services
If your target is not listed, we offer custom PNA design and synthesis. Whether you require sequence optimization, specific modifications, or larger production quantities, our team can support your project. Submit your project details and we will provide a personalized proposal.
References
Clinical Biochemistry Volume 40, Issue 7, April 2007, Pages 499-502
Functional identity of genes detectable in expression profiling assays following globin mRNA reduction of peripheral blood samples
Objectives:
Design and methods:
Results:
Conclusions:
2016 Aug 12:6:31584. doi: 10.1038/srep31584.
Globin mRNA reduction for whole-blood transcriptome sequencing
The transcriptome analysis of whole-blood RNA by sequencing holds promise for the identification and tracking of biomarkers; however, the high globin mRNA (gmRNA) content of erythrocytes hampers whole-blood and buffy coat analyses. We introduce a novel gmRNA locking assay (GlobinLock, GL) as a robust and simple gmRNA reduction tool to preserve RNA quality, save time and cost. GL consists of a pair of gmRNA-specific oligonucleotides in RNA initial denaturation buffer that is effective immediately after RNA denaturation and adds only ten minutes of incubation to the whole cDNA synthesis procedure when compared to non-blood RNA analysis. We show that GL is fully effective not only for human samples but also for mouse and rat, and so far incompletely studied cow, dog and zebrafish.
29 Jan 2008 - https://doi.org/10.2217/17520363.2.1.11
Comparison of Globin RNA Processing Methods for Genome-Wide Transcriptome Analysis from Whole Blood
