Fungal ITS2 PNA
Fungal ITS2 PCR Blocker
Fungal ITS2 is a high-performance PNA oligonucleotide engineered to selectively inhibit the amplification of the ITS2 rRNA region. In fungal genomic studies, ITS2 often acts as an overrepresented signal that masks more valuable genetic data. By specifically binding to the conserved cgagggcacgtctgcctgg sequence, this blocker silences dominant ITS signals, allowing researchers to uncover rare nuclear genes and low-abundance targets that would otherwise remain undetected.
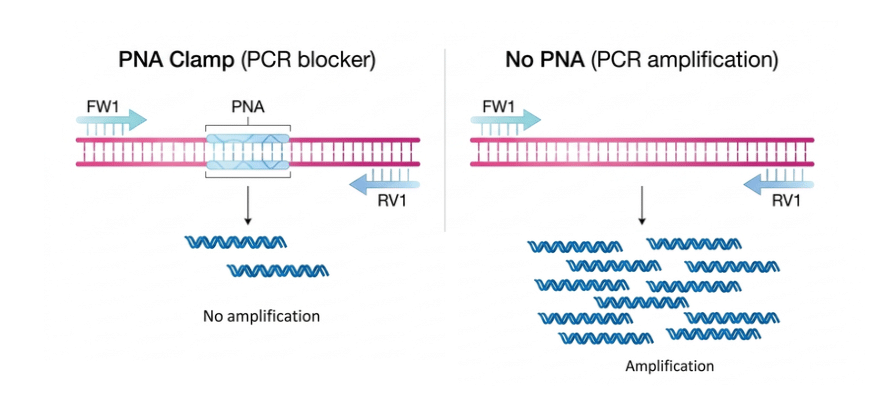
Main characteristics
Fungal ITS2 PNA selectively binds conserved fungal ITS2 rRNA sequences, preventing their amplification in ITS-based assays. Thanks to its stable PNA backbone, it is resistant to nuclease degradation and ensures high specificity. This reduces off-target amplification and improves the accuracy of PCR and metagenomic workflows, making it a reliable tool for complex DNA analysis. A terminal glycine is added to the PNA structure to improve flexibility and reduce steric hindrance, which can help optimize binding efficiency without affecting specificity.
Applications
- Suppresses fungal ITS2 rRNA during PCR and qPCR for clearer DNA/RNA detection
- Enables accurate fungal profiling in microbiome studies
- Supports biodiversity and ecological metagenomic research
- Suitable for environmental and clinical sample analysis
- Improves NGS results through effective rRNA depletion
Technical specification
 |
Sequency : | cgagggcacgtctgcctgg-G |
 |
MW : | 5513.18g/mol |
 |
Purity : | > 95% |
 |
Counter-Ion : | TFA Salts |
 |
Delivery format : | Lyophilized |
Price
| Product | Size | Price € |
Price $ |
| SB335 - 25nmol | 25 nmol | 433 | 520 |
| SB335 - 50 nmol | 50 nmol | 705 | 846 |
For more information about PNA, please visit our dedicated page.
Explore our PNA Catalog products. Click HERE to discover additional reference compounds.
Custom PNA Services
If your target is not listed, we offer custom PNA design and synthesis. Whether you require sequence optimization, specific modifications, or larger production quantities, our team can support your project. Submit your project details and we will provide a personalized proposal.
References
2024 Feb 4;15(2):255-271. doi: 10.1080/21501203.2023.2301003. eCollection 2024.
Primer pairs, PCR conditions, and peptide nucleic acid clamps affect fungal diversity assessment from plant root tissues
Abstract
High-throughput sequencing has become a prominent tool to assess plant-associated microbial diversity. Still, some technical challenges remain in characterising these communities, notably due to plant and fungal DNA co-amplification. Fungal-specific primers, Peptide Nucleic Acid (PNA) clamps, or adjusting PCR conditions are approaches to limit plant DNA contamination. However, a systematic comparison of these factors and their interactions, which could limit plant DNA contamination in the study of plant mycobiota, is still lacking. Here, three primers targeting the ITS2 region were evaluated alone or in combination with PNA clamps both on nettle (Urtica dioica) root DNA and a mock community. PNA clamps did not improve the richness or diversity of the fungal communities but increased the number of fungal reads. Among the tested factors, the most significant was the primer pair. Specifically, the 5.8S-Fun/ITS4-Fun pair exhibited a higher OTU richness but fewer fungal reads. Our study demonstrates that the choice of primers is critical for limiting plant and fungal DNA co-amplification. PNA clamps increase the number of fungal reads when ITS2 is targeted but do not result in higher fungal diversity recovery at high sequencing depth. At lower read depths, PNA clamps might enhance microbial diversity quantification for primer pairs lacking fungal specificity.
01 February 2021 https://doi.org/10.1111/nph.17248
Fire alters plant microbiome assembly patterns: integrating the plant and soil microbial response to disturbance
Abstract
- It is increasingly evident that the plant microbiome is a strong determinant of plant health. While the ability to manipulate the microbiome in plants and ecosystems recovering from disturbance may be useful, our understanding of the plant microbiome in regenerating plant communities is currently limited.
- Using 16S ribosomal RNA (rRNA) gene and internal transcribed spacer (ITS) region amplicon sequencing, we characterized the leaf, stem, fine root, rhizome, and rhizosphere microbiome of < 1-yr-old aspen saplings and the associated bulk soil after a recent high-intensity prescribed fire across a burn severity gradient.
- Consistent with previous studies, we found that soil microbiomes are responsive to fire. We extend these findings by showing that certain plant tissue microbiomes also change in response to fire. Differences in soil microbiome compositions could be attributed to soil chemical characteristics, but, generally, plant tissue microbiomes were not related to plant tissue elemental concentrations. Using source tracking modeling, we also show that fire influences the relative dominance of microbial inoculum and the vertical inheritance of the sapling microbiome from the parent tree.
- Overall, our results demonstrate how fire impacts plant microbiome assembly, diversity, and composition and highlights potential for further research towards increasing plant fitness and ecosystem recovery after fire events.
Research Article 18 July 2022 - https://doi.org/10.1128/msystems.00120-22
Relationships between Sphaerulina musiva Infection and the Populus Microbiome and Metabolome
Abstract
Pathogenic fungal infections in plants may, in some cases, lead to downstream systematic impacts on the plant metabolome and microbiome that may either alleviate or exacerbate the effects of the fungal pathogen. While Sphaerulina musiva is a well-characterized fungal pathogen which infects Populus tree species, an important wood fiber and biofuel feedstock, little is known about its systematic effects on the metabolome and microbiome of Populus. Here, we investigated the metabolome of Populus trichocarpa and Populus deltoides leaves and roots and the microbiome of the leaf and root endospheres, phylloplane, and rhizosphere to understand the systematic impacts of S. musiva abundance and infection on Populus species in a common garden field setting. We found that S. musiva is indeed present in both P. deltoides and P. trichocarpa, but S. musiva abundance was not statistically related to stem canker onset. We also found that the leaf and root metabolomes significantly differ between the two Populus species and that certain leaf metabolites, particularly the phenolic glycosides salirepin and salireposide, are diminished in canker-infected P. trichocarpa trees compared to their uninfected counterparts. Furthermore, we found significant associations between the metabolome, S. musiva abundance, and microbiome composition and α-diversity, particularly in P. trichocarpa leaves. Our results show that S. musiva colonizes both resistant and susceptible hosts and that the effects of S. musiva on susceptible trees are not confined to the site of canker infection.
24 May 2023 https://doi.org/10.1111/1758-2229.13167
The ‘microbiome counterattack’: Insights on the soil and root-associated microbiome in diverse chickpea and lentil genotypes after an erratic rainfall event
Abstract
Legumes maintain soil fertility thanks to their associated microbiota but are threatened by climate change that causes soil microbial community structural and functional modifications. The core microbiome associated with different chickpea and lentil genotypes was described after an unexpected climatic event. Results showed that chickpea and lentil bulk soil microbiomes varied significantly between two sampling time points, the first immediately after the rainfall and the second 2 weeks later. Rhizobia were associated with the soil of the more productive chickpea genotypes in terms of flower and fruit number. The root-associated bacteria and fungi were surveyed in lentil genotypes, considering that several parcels showed disease symptoms. The metabarcoding analysis revealed that reads related to fungal pathogens were significantly associated with one lentil genotype. A lentil core prokaryotic community common to all genotypes was identified as well as a genotype-specific one. A higher number of specific bacterial taxa and an enhanced tolerance to fungal diseases characterized a lentil landrace compared to the commercial varieties. This outcome supported the hypothesis that locally adapted landraces might have a high recruiting efficiency of beneficial soil microbes.
